GENIOSave is Geniomic’s solution for high-efficiency genomic data management and storage:
| Lossless compression reducing FASTQ.gz files by 30% and sorted BAM files by 50%, on average (see our benchmark) |
A versatile genomic data compression toolkit especially suited to tabular formats such as FASTQ, FASTQ.gz, SAM, BAM and other file formats as well. |
| Direct access to your target genomic data by real-time decompression |
Direct use of compressed files via pipe-enabled applications. No need for further format conversion. A single file format for many applications dramatically reduces your storage needs. |
| Enhanced algorithm allowing selective access to compressed genomic data |
Whatever the origin of your file, embedded indexing guarantees selective access to chromosome regions. |
GENIOSave: reduce your genomic costs!
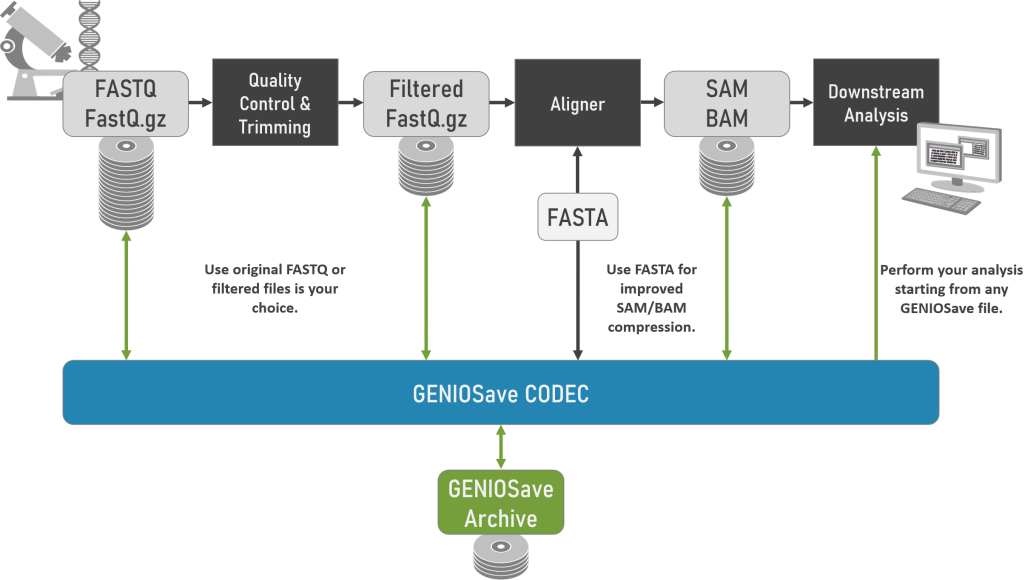
Highlights
- The GENIOSave technology compresses tabular data formats optimised for genomic files.
- The compressed files are organised according to addressable genomic regions.
- The decoder can selectively decode genomic regions thanks to an embedded index.
- The encoder automatically recognizes the input file format producing the compressed file.
- The compressed file losselessly retains all the of the information in the original file, including the optional fields.
- Improved encoding using a reference genome.
- The decoder produces the user-specified file format.
Main features
- Multiple format compression: different genomic file formats can be reversibly compressed to a single GENIOSave file.
- Piping: possibility not to store genomic files used in downstream analysis.
- Accelerated decoding: decoding restricted to user-selected fields, e.g., sequence_ID, quality values, etc.
- Embedded sorting: GENIOSave file records can be ordered by position during encoding.
- Embedded access index: a GENIOSave file includes an index for selective access.
- User-selectable compressed block size: selectable size of a compressed block of reads.
- Compression without reference genome: if the MD field is present, reference genome is not required.
- Selective access: user can select a target genomic region. GENIOSave decodes only the requested compressed data.

